ISSN: 0970-938X (Print) | 0976-1683 (Electronic)
Biomedical Research
An International Journal of Medical Sciences
Research Article - Biomedical Research (2018) Volume 29, Issue 5
FGFR1 amplification and FGFR gene fusion in resected squamous cell carcinoma of the lung
Lei Wang1#, Juan Liu2#, Mingsong Wang1, Haibo Xiao1, Guoqing Li1, Fengqing Hu1* and Ju Mei1*
1Department of Cardiothoracic Surgery, Xin Hua Hospital Affiliated to Shanghai Jiao Tong University School of Medicine, Shanghai, 200092, PR China
2Department of Nursing, Baoshan Branch of Shanghai Shuguang Hospital, Shanghai University of Traditional Chinese Medicine, Shanghai, 201900, PR China
#These authors contributed to this work equally and should be considered co-first authors
- *Corresponding Authors:
- Fengqing Hu
Department of Cardiothoracic Surgery
Xin Hua Hospital Affiliated to Shanghai Jiao Tong University School of Medicine, PR China
Ju Mei
Department of Cardiothoracic Surgery
Xin Hua Hospital Affiliated to Shanghai Jiao Tong University School of Medicine, PR China
Accepted date: November 28, 2017
DOI: 10.4066/biomedicalresearch.29-17-3086
Visit for more related articles at Biomedical ResearchThe objective of the present study was to investigate the clinicopathologic characteristics and prognostic role of FGFR1 amplification and FGFR fusion in patients with surgically resected squamous cell carcinoma of the lung (SCCL). Here, fluorescent in situ hybridization (FISH) was performed to detect FGFR1 amplification and reverse transcriptase polymerase chain reaction (RT-PCR) was used to screen 15 known FGFR fusion variants in 108 patients with surgically resected SCCL. All cases were also analyzed for EGFR, KRAS, HER2 and BRAF mutations. Clinical characteristics including age, sex, smoking status, stage, relapse-free survival (RFS) and overall survival (OS) were collected. Of 108 tumors screened, 14 (13.0%) FGFR1 amplification was found. There were 4 (3.7%) patients that harbored FGFR fusion including 2 BAG4-FGFR1 and 2 FGFR3-TACC3 fusion. Compared to the FGFR1 amplification negative group, patients with FGFR1 amplification were more likely to be smokers (100.0%, 14 of 14 patients, p=0.036), significantly associated with larger tumor (>3 cm) (88.2%, 13 of 14 patients, p=0.032). Patients with FGFR1 amplification had worse RFS (p=0.013) and OS (p=0.021) than those without FGFR1 amplification. There was no correlation between FGFR fusion and clinicopathologic characteristics. No significant difference in RFS or OS was found between patients with FGFR fusion and those without FGFR fusion. In conclusion, FGFR1 amplification and FGFR fusion occurred in 13.0% and 3.7% of patients with surgically resected SCCL, separately. FGFR1 amplification was correlated with poor prognosis and identified a distinct subset of SCCL with a higher prevalence among smokers with relative larger tumor (>3 cm). FGFR is a therapeutic target and patients with FGFR1 amplification or FGFR fusion may benefit from FGFR targeted therapy which needs further clinical investigation.
Keywords
FGFR1 amplification, FGFR fusion, Squamous cell carcinoma of the lung
Introduction
Lung cancer is the leading cause of cancer mortality worldwide resulting in more than 1 million deaths annually [1-3]. Because the activating mutation in the epidermal growth factor receptor (EGFR) is associated with dramatic responses to EGFR tyrosine kinase inhibitors, and the oncogenic fusion genes such as ALK, ROS1 or RET rearrangement are related to sensitivity of treatment with crizotinib or vandetanib, identification of oncogenic driver genes and molecularly targeted therapies based on tumor biology in NSCLC are becoming increasingly important [4-7]. However, most of advances in personalized treatment was about adenocarcinoma of lung and effective targeted therapies for squamous cell carcinoma of the lung (SCCL) [6,8], accounting for approximately 25% of non-smallcell lung cancers (NSCLCs), remained elusive. As a result, it will be necessary to clarify different molecular alteration on SCCL for further individual treatment.
The FGFR family, comprising FGFR1, FGFR2, FGFR3 and FGFR4, play crucial roles in cancer development and are targets for dysregulation by amplification, point mutations, or translocation [5,9,10]. Amplified FGFR1 has been reported in 20% of SCCL and inhibition of the FGFR1 pathway with FGFR inhibitors was demonstrated to lead to significant tumor shrinkage, suggesting that FGFR inhibitors might be an effective therapeutic option in SCCL with FGFR1 amplification [11,12]. Recently, FGFR fusion was identified in diverse cancers including glioblastoma, bladder cancer, head and neck cancer and SCCL [13,14]. Moreover, FGFR fusion has been shown to sensitize cancer cells to FGFR kinase inhibitors, suggesting that a new subset of cancers maybe treatable with FGFR targeted therapy [13].
To date, there are few studies that constitute a comprehensive picture of FGFR1 amplification, FGFR fusion, their protein expression and correlation to SCCL [15]. In the present study, we examined FGFR1 gene amplification, FGFR gene fusion, as well as the expression of FGFR1/FGFR3 in a consecutive collection of SCCL tumor samples. The mutational status of known oncogenic genes, including EGFR, KRAS, HER2 and BRAF were also examined. This detailed understanding of FGFR alterations and other oncogene mutations in SCCL might enable a more precise delineation of candidate target populations, facilitating clinical trial design and validation of predictive biomarkers.
Materials and Methods
Patients and tissues
From October 2011 to September 2013, we consecutively procured primary tumor samples and corresponding formalinfixed, paraffin-embedded tumor blocks from SCCL patients who underwent pulmonary resection at the Department of Cardiothoracic Surgery, Xinhua Hospital, Shanghai Jiaotong University school of medicine. Subjects eligible for this study had to meet the following: confirmed SCCL diagnosis by hematoxylin-eosin (H&E) and IHC staining, each sample containing sufficient for comprehensive mutational analyses and with no neoadjuvant treatment. This research was approved by the Institutional Review Board of the Xinhua Hospital, Shanghai Jiaotong University School of medicine. Written informed consent from each patient was obtained. Clinical and pathologic data were prospectively collected for analyses including age at diagnosis, sex, smoking history, histologic type, pathologic TNM stage, tumor size, tumor differentiation. Patients were observed in clinic or by telephone for disease recurrence and survival from the date of diagnosis.
Reverse Transcription Polymerase Chain Reaction (RT-PCR) and FGFR fusions
Frozen tumor specimens were dissected and RNA/DNA was co-extracted following the standard instructions of RNA/DNA isolation Kit (TIANGEN BIOTECH, Beijing, China). Singlestranded RNA of each sample is reverse transcribed into complementary DNA (cDNA) by RevertAid First Strand cDNA Synthesis Kit (Fermentas, EU). For detection of FGFR fusions, primers were designed to cover all known fusion variants. Direct sequencing in both forwards and reverse directions was then performed in PCR amplification products. Details of RT-PCR are provided in the appendix.
Mutational analyses
EGFR (exons 18-21), KRAS (exons 2-3), HER2 (exons 18-21), BRAF (exons 11-15) were amplified with KOD Plus Neo DNA polymerase (Toyobo, Osaka, Japan). All mutated cases were confirmed twice with independent PCR reactions.
Assessment of FGFR1 gene amplification
Fluorescent in situ hybridization (FISH) assay for FGFR1 was performed by using FGFR1 probe that hybridizes to the band 8p12-8p11.23 with Spectrum Orange (red) CEP8 with Spectrum Green (Abbott Molecular, Abbott Park, IL) following routine methods. FISH analysis was reinterpreted by two experienced evaluators blinded to the clinical data. At least 100 nuclei per patient were evaluated. Tissue samples with a FGFR1/CEP8 ratio of 1.0 were classified as normal and those with a FGFR1/CEP8 ratio between 1.0 and 2.0 were classified as having FGFR1 gains. A FGFR1/CEN8 ratio of more than 4.0 was considered amplified. A minimum of 50 cells with both centromeric and FGFR1 gene signals were scored to give conclusive data.
Immunohistochemistry analysis
To evaluate immunohistochemistry (IHC) analysis for screening and detecting SCCL with FGFR fusion genes, all of the included samples were subjected to IHC analysis with formalin-fixed paraffin-embedded slides sectioned. Details of IHC method is provided in the Appendix.
Results
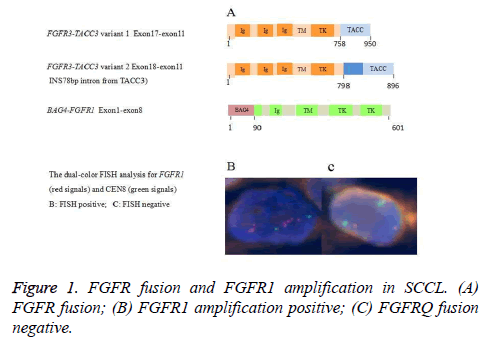
FGFR1 amplification of SCCL by fluorescence in situ hybridization
A total of 108 cases met eligibility for this study. The characteristics for these patients are listed in Appendix Table A1. We analyzed the enrolled cases with gene copy number equal or above 4, a threshold that has been previously used to define FGFR1 amplifications [12]. Among the 108 patients, FGFR1 amplification was found in 13.0/% (14/108) of the cases (Figure 1). Details of the 14 cases with FGFR1 amplification are listed in Table 1. The frequency of FGFR1 amplification was significantly higher in current or ever smokers than in never-smokers (100% vs. 0%; p=0.036). Patients with FGFR1 amplification were also significantly associated with larger tumor (>3 cm) (92.9%, 13 of 14 patients, p=0.032). However, There was no significant association between increased FGFR1 gene copy number (≥ 4) and other clinicopathological parameter (sex, p=0.593; age, p=0.577; stage, p=1.000; tumor differentiation, p=1.000) (Table 2).
| Patient No. | Age (years) | Sex | Smoking (pack-years) | Stage | Tumor Size (cm) | Lymph Node Status | Differentiation | FGFR1 FISH | FGFR3 fusion |
|---|---|---|---|---|---|---|---|---|---|
| 1 | 58 | M | 100 | IIa | 7.0 | 0 | Moderate | amp | N |
| 2 | 51 | M | 30 | IIa | 4.5 | N1 | Moderate | amp | N |
| 3 | 57 | M | 100 | IIIa | 4.0 | N2 | Poor | amp | N |
| 4 | 75 | M | 60 | Ib | 1.4 | 0 | Poor | amp | N |
| 5 | 64 | M | 80 | IIb | 9.0 | 0 | Poor | amp | N |
| 6 | 72 | M | 35 | IIIa | 5.2 | N2 | Poor | amp | N |
| 7 | 64 | M | 120 | IIa | 5.5 | 0 | Moderate | amp | N |
| 8 | 66 | M | 40 | Ib | 4.0 | 0 | Moderate | amp | N |
| 9 | 64 | M | 40 | IIa | 4.0 | N1 | Poor | amp | N |
| 10 | 45 | M | 38 | IIIa | 4.8 | N2 | Poor | amp | N |
| 11 | 58 | M | 30 | IIa | 5.0 | N1 | Poor | amp | N |
| 12 | 62 | M | 15 | Ib | 5.0 | 0 | Moderate | amp | N |
| 13 | 63 | M | 50 | IIb | 10.0 | 0 | Poor | amp | N |
| 14 | 61 | M | 160 | Ib | 3.5 | 0 | Moderate | amp | N |
| 15 | 49 | M | 20 | IIIa | 4.0 | N2 | Moderate | N | FGFR3-TACC3Δ |
| 16 | 65 | M | 40 | Ib | 5.0 | 0 | Poor | N | FGFR3-TACC3* |
| 17 | 61 | M | 40 | IIb | 5.5 | 0 | Moderate | N | BAG4--FGFR1 |
| 18 | 48 | M | 105 | Ib | 4 | 0 | Poor | N | BAG4--FGFR1 |
Table 1. Clinicopathologic Details of 18 SCCL Patients with FGFR1 amplification or FGFR3 Fusions.
| Characteristic | FGFR1 Amp (+) | FGFR1 Amp (-) | P |
|---|---|---|---|
| No.of patients (%) | No.of patients (%) | ||
| Age, years | |||
| ≤ 60 | 5 (35.7) | 42 (44.7) | 0.577 |
| >60 | 9 (64.3) | 52 (55.3) | |
| Sex | |||
| Male | 14 (100.0) | 86 (91.5) | 0.593 |
| Female | 0 (0.0) | 8 (8.5) | |
| Smoking Status | |||
| N | 0 (0.0) | 24 (25.5) | 0.036 |
| C/E | 14 (100.0) | 70 (74.5) | |
| Tumor size | |||
| ≤ 3 cm | 1 (7.1) | 36 (38.3) | 0.032 |
| >3 cm | 13 (92.9) | 58 (61.7) | |
| Stage | |||
| I~II | 11 (78.6) | 71 (75.6) | 1 |
| III~IV | 3 (21.4) | 23 (24.5) | |
| Differentiation | |||
| Well-moderate | 6 (42.9) | 40 (42.6) | 1 |
| Poor | 8 (57.1) | 54 (57.4) | |
Table 2. Clinicopathologic characteristics of FGFR1 amplification in SCCL.
Frequency and clinicopathologic characteristics of FGFR fusion in SCCL
All tumors were examined by RT-PCR and direct sequencing with primer sets covered 15 known FGFR fusion variants. Four FGFR fusions in 108 SCCL were identified. Of 108 squamous cell carcinomas, 4 (3.7%) harbored FGFR fusions which included two BAG4-FGFR1 fusions and two FGFR3-TACC3 fusions. Details of the 4 FGFR fusion positive SCCLs are listed in Table 1. The histology of 4 tumors with FGFR fusions were confirmed by IHC bio-markers (p40 and TTF-1). We found that all of 4 FGFR fusion positive patients were current or ever smokers with lager tumor (>3 cm) while the frequency of was considerably higher than those in never smokers with tumors which were less than 3cm. This difference, however, failed to reach statistical significance (p=0.573 in smoking status and p=0.297 in tumor size). Similarly, there was no significant association between FGFR fusion and clinicopathological parameter (sex, p=1.000; age, p=1.000; stage, p=1.000; tumor differentiation, p=1.000) (Appendix Table A2).
Mutation analysis and IHC analysis
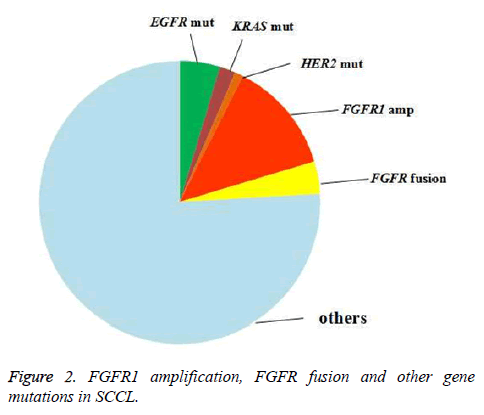
To explore the relationship of FGFR1 amplification/FGFR fusion with other oncogene mutations in SCCL, testing for EGFR, KRAS, HER2 and BRAF gene mutation was also arranged. Twenty-six patients were found to harbor EGFR, HER2, KRAS or BRAF mutation, including 4.6% (5/108) with an EGFR mutation, 1.9% (2/108) with a KRAS mutation and 0.9% (1/108) with an HER2 mutation. BRAF mutation was not found in our study (Figure 2). All of the FGFR1 amplification, FGFR fusion and oncogene mutation that we tested were excluded. Neither FGFR1 amplification nor FGFR fusion was found to be correlated with the oncogene mutations above (Appendix Tables A3 and A4).
Furthermore, we detected the relationship between FGFR1 amplification and FGFR fusion by IHC. Nevertheless, no significant correlation was found, either. (Appendix Table A5).
Clinical outcome
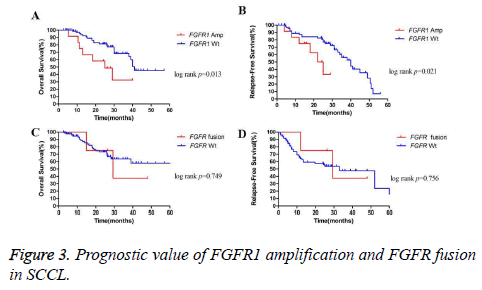
The overall survival (OS) for all of the patients was 25.0 months for a median follow-up time of 25.2 months. As expected, median OS was longer in patients with stage I disease than in those with stage II to III disease (39.0 months vs. 19.5 months; p=0.011). A similar trend was also found in OS between SCCL patients without or with lymph node metastasis (26.2 months vs. 19.0 months; p=0.056). We did not observe significant differences in OS according to other clinicopathological parameters such as age, sex, smoking status or tumor tumor grade (p=0.261 for age, p=0.553 for sex, p=0.553 for sex, p=0.322 for smoking status and p=0.937 for tumor grade). We further detect the prognostic value of FGFR1 gene amplification and FGFR fusion in the same panel of 108 cases. Survival for patients with FGFR1 amplification was shorter than patients without amplification (25.0 months vs. 41.0 months; p=0.013) (Figure 3A). However, no significant difference in OS was found between patients with or without FGFR fusion (27.8 months vs. 25.0 months; p=0.749) (Figure 3C).
The median recurrence-free survival (RFS) for the cohort was 15.8 months. Like OS, median RFS was longer in patients with stage I disease than in those with stage II to III disease (52.0 months vs. 12.7 months; p=0.029). A significant difference in RFS was also found between patients with or without lymph node metastasis (50.0 months for patients without lymph node metastasis vs. 11.7 months for patients with lymph node metastasis, p=0.040). Similar to OS, FGFR1 amplified cases harbored worse RFS than those FGFR1 wild-type patients (23.5 months vs. 40.0 months; p=0.021) (Figure 3B). Nevertheless, the difference in RFS between patients with or without FGFR fusion was not significant (29.3 months for patients with FGFR fusion vs. 33.0 months for patients without FGFR fusion, p=0.756) (Figure 3D).
Discussion
FGFR1 amplification and FGFR fusion seem to be of key importance in tumor progression and were illustrated to be new drivers for a range of cancers [16,17]. It was reported that FGFR1 amplification was found in about 10% of breast cancer and located on chromosome 8p12 [18]. Other tumors, including bladder cancer, ovarian carcinoma and rhabdomyosarcoma, were also tested and FGFR1 amplification were found with lower frequency at the same region [19]. Recently, it is shown that gene rearrangement also contributes to the developments of solid tumors and the FGFR fusion gene was illustrated to play important roles in divers cancers [13]. The oncogenic potential of FGFR1-TACC1, FGFR2-BICC1 and FGFR3-TACC3 were confirmed by expression of the fusion kinases in NIH3T3 fibroblasts, Rat1A fibroblasts or astrocytes and conferred in vivo tumorigenesis of subcutaneous transplanted cells in immune-deficient mice [18,20].
In our study, we investigated whether FGFR1 amplification and FGFR fusion were associated with the clinicopathologic parameters and their impact on survival in patients with operable SCCL. To the best of our knowledge, this may be the first report of multiple analysis on FGFR1 amplification, FGFR fusion and oncogene mutations in a cohort of East Asians with SCCL, and we illustrated two important subtypes, showing negative prognostic impact of FGFR1 amplification and demonstrating FGFR fusion as a unique subtype in resectable SCCL.
We demonstrated that patients with FGFR1 amplification had identifiable clinicopathologic characteristics, including smokers status and relative larger tumor (>3 cm). In principle, our analysis showed similar results as shown by Weiss et al. They found that increased copy number status of FGFR1 (≥ 4 signals) in lung SCC correlated significantly with the gender, smoking status and pathological subtypes but not with age, grade and lymph node status [12]. Interestingly, tumor size might be firstly recognized to be related with FGFR1 amplification in our research. However, Lukas et al. reported no significant associations between FGFR1 amplification and clinicopathological parameters [17]. The lack of standardized criteria for defining FGFR1 amplification and the different number of cases may be the possible reason [12,17].
Another interesting founding in our research is that patients with FGFR1 amplification had significantly shorter RFS and OS than those without FGFR1 amplification. This result suggests that targeting the FGFR1 pathway might improve survival in patients with FGFR1 amplification who have resectable SCCL. Nowadays, several potent FGFR tyrosine kinase inhibitors including BGJ398 and AZD4547 have been in early clinical research [21,22]. However, there are more gene alterations that were found to be correlated with prognosis in adenocarcinoma than in SCCL. More comprehensive analysis will be needed to explore how to precisely predict the prognosis of patients with SCCL.
In the current study, we detected the FGFR1, FGFR2, FGFR3 gene fusions in SCCL patients and found the frequency of FGFR fusions was 3.7%, which is similar to previous studies [9]. FGFR fusions and FGFR1 amplification were mutually exclusive with activating mutations in EGFR, KRAS, BRAF and HER2, showing the driver’s role of those gene alteration in cancer development. Although a variety of fusion partners of FGFR were reported by recent studies [23,24], only one partner each for the FGFR1 and FGFR3 fusions (BAG4- FGFR1 and FGFR3-TACC3) were found. FGFR2 fusions were not identified in our study, indicating that FGFR3 or FGFR1, other than FGFR2 might be crucial for oncogenesis in SCCL. Unlike the previous study, we failed to find the correlation between FGFR gene fusion and clinicopathologic characteristics. This may be duo to different pathology subtypes of the cohort (the precious study showed the correlation in NSCLC, but we detected that in SCCL). The slightly lower number of cases that were analysed in our study may be another resean.
The methods for detection of gene fusions include RT-PCR, IHC, and FISH. Nevertheless, FISH assay was not available because the interval of two genes (FGFR3-TACC3 or BAG4- FGFR1) was too small to detect rearrangement. In current study, 14 known fusion variants of FGFR found in glioblastoma, cholangiocarcinoma, bladder cancer, thyroid cancer, oral cancer, breast cancer or head and neck squamous cell cancer by previous study were tested in 108 SCCLs [14]. Since RT-PCR could not be used to detect unknown fusion variants, it is possible that a few FGFR rearranged samples may have been missed. Therefore, more studies will be warranted to validate our findings in an independent cohort, especially in an Eastern population.
In conclusion, our data demonstrated that FGFR1 amplification and FGFR fusion occurred in 13.0% and 3.7% of patients with lung squamous cell carcinoma, separately. FGFR1 amplification was correlated with poor prognosis and identified a distinct subset of SCCL with a higher prevalence among smokers with relative larger tumor (>3 cm). Knowledge of these clinical characteristics will help clinicians select those patients most likely to harbor this genetic alteration and most likely to benefit from FGFR targeted therapy.
Acknowledgements
This study was supported by Shanghai Sailing Program for Young Scientific and Technological Talents (15YF1407800) and National Natural Science Foundation of China (NSFC) 81602418 a 81572248.
References
- Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA: J Clin 2015; 65: 87-108.
- Chia PL, John T. Severe psoriasis flare after anti-programmed death ligand 1 (PD-L1) therapy for metastatic non-small cell lung cancer (NSCLC). J Immunother 2016; 39: 202-204.
- Printz C. Lung cancer new leading cause of death for women in developed countries: Data reflects increased rates of smoking. Cancer 2015; 121: 1911-1912.
- Bergethon K, Shaw AT, Ou SH, Katayama R, Lovly CM, McDonald NT. ROS1 rearrangements define a unique molecular class of lung cancers. J Clin Oncol 2012; 30: 863-870.
- Kohno T, Tsuta K, Tsuchihara K, Nakaoku T, Yoh K, Goto K. RET fusion gene: translation to personalized lung cancer therapy. Cancer Sci 2013; 104: 1396-400.
- Dahabreh IJ, Linardou H, Siannis F, Kosmidis P, Bafaloukos D, Murray S. Somatic EGFR mutation and gene copy gain as predictive biomarkers for response to tyrosine kinase inhibitors in non-small cell lung cancer. Clin Cancer Res 2010; 16: 291-303.
- Miyanaga A, Shimizu K, Noro R, Seike M, Kitamura K, Kosaihira S. Activity of EGFR-tyrosine kinase and ALK inhibitors for EML4-ALK-rearranged non-small-cell lung cancer harbored coexisting EGFR mutation. BMC Cancer 2013; 13: 262.
- Sun Y, Ren Y, Fang Z, Li C, Fang R, Gao B. Lung adenocarcinoma from East Asian never-smokers is a disease largely defined by targetable oncogenic mutant kinases. J Clin Oncol 2010; 28: 4616-4620.
- Hibi M, Kaneda H, Tanizaki J, Sakai K, Togashi Y, Terashima M. FGFR gene alterations in lung squamous cell carcinoma are potential targets for the multikinase inhibitor nintedanib. Cancer Sci 2016; 107: 1667-1676.
- Wang R, Wang L, Li Y, Hu H, Shen L, Shen X. FGFR1/3 tyrosine kinase fusions define a unique molecular subtype of non-small cell lung cancer. Clin Cancer Res 2014; 20: 4107-4114.
- Dutt A, Ramos AH, Hammerman PS, Mermel C, Cho J, Sharifnia T. Inhibitor-sensitive FGFR1 amplification in human non-small cell lung cancer. PloS one 2011; 6: e20351.
- Weiss J, Sos ML, Seidel D, Peifer M, Zander T, Heuckmann JM. Frequent and focal FGFR1 amplification associates with therapeutically tractable FGFR1 dependency in squamous cell lung cancer. Sci Translat Med 2010; 2: 62-93.
- Sabnis AJ, Bivona TG. FGFR fusions in the driver's seat. Cancer Discov 2013; 3: 607-609.
- Wu YM, Su F, Kalyana-Sundaram S, Khazanov N, Ateeq B, Cao X. Identification of targetable FGFR gene fusions in diverse cancers. Cancer Discov 2013; 3: 636-647.
- Zheng S, Pan Y, Wang R, Li Y, Cheng C, Shen X. SOX2 expression is associated with FGFR fusion genes and predicts favorable outcome in lung squamous cell carcinomas. OncoTargets Ther 2015; 8: 3009-3016.
- Carneiro BA, Elvin JA, Kamath SD, Ali SM, Paintal AS, Restrepo A. FGFR3-TACC3: A novel gene fusion in cervical cancer. Gynecol Oncol Rep 2015; 13: 53-56.
- Kohler LH, Mireskandari M, Knosel T, Altendorf-Hofmann A, Kunze A, Schmidt A. FGFR1 expression and gene copy numbers in human lung cancer. Virchows Arch 2012; 461: 49-57.
- Marshall ME, Hinz TK, Kono SA, Singleton KR, Bichon B, Ware KE. Fibroblast growth factor receptors are components of autocrine signaling networks in head and neck squamous cell carcinoma cells. Clinical Cancer Res 2011; 17: 5016-5025.
- Kim HR, Kim DJ, Kang DR, Lee JG, Lim SM, Lee CY. Fibroblast growth factor receptor 1 gene amplification is associated with poor survival and cigarette smoking dosage in patients with resected squamous cell lung cancer. J Clin Oncol 2013; 31: 731-737.
- Kim SJ, Kim HG, Lim HW, Park EH, Lim CJ. Up-regulation of glutathione biosynthesis in NIH3T3 cells transformed with the ETV6-NTRK3 gene fusion. Mol Cell 2005; 19: 131-136.
- Allison KH, Dintzis SM, Schmidt RA. Frequency of HER2 heterogeneity by fluorescence in situ hybridization according to CAP expert panel recommendations: time for a new look at how to report heterogeneity. Am J Clin Pathol 2011; 136: 864-871.
- Pandiar D, Shameena P. Immunohistochemical expression of CD34 and basic fibroblast growth factor (bFGF) in oral submucous fibrosis. J Oral Maxillofacial Pathol 2014; 18: 155-161.
- Arai Y, Totoki Y, Hosoda F, Shirota T, Hama N, Nakamura H. Fibroblast growth factor receptor 2 tyrosine kinase fusions define a unique molecular subtype of cholangiocarcinoma. Hepatology 2014; 59: 1427-1434.
- Petersen I, Kotb WF, Friedrich KH, Schluns K, Bocking A, Dietel M. Core classification of lung cancer: correlating nuclear size and mitoses with ploidy and clinicopathological parameters. Lung Cancer 2009; 65: 312-318.