ISSN: 0970-938X (Print) | 0976-1683 (Electronic)
Biomedical Research
An International Journal of Medical Sciences
Research Article - Biomedical Research (2017) Volume 28, Issue 7
In vitro antifungal effects of biosynthesized silver nanoparticle by Candida albicans against Candida glabrata
Iman Bahrami Abdehgah1, Alireza Khodavandi2, Ali Shamsazar3, Masoud Negahdary4, Mahnaz Jafarzadeh5 and Ghasem Rahimi6*
1Young Researchers and Elite Club, Yasooj Branch, Islamic Azad University, Yasooj, Iran
2Department of Biology, Gachsaran Branch, Islamic Azad University, Gachsaran, Iran
3Department of Biochemistry, Payame Noor University, Tehran, Iran
4Yazd Cardiovascular Research Center, Shahid Sadoughi University of Medical Sciences, Yazd, Iran
5Department of Physics, Faculty of Sciences, Urmia University, P.B. 165, Urmia, Iran
6Young Researchers and Elite Club, Marvdasht Branch, Islamic Azad University, Marvdasht, Iran
Accepted date: July 12, 2016
Recently, biosynthesis of silver nanoparticles (NPs), due to low toxicity and high compatibility with the environment, has added to the antimicrobial properties of this substance. In this study, silver NPs were synthesized from Candida albicans and their antifungal effects against Candida glabrata were evaluated. First, silver NPs using clinical strains of Candida albicans were synthesized in the presence of silver nitrate 1.5 mM at room temperature. Then, physicochemical properties of the synthesized nanoparticles were explored using Scanning Electron Microscope (SEM), Ultraviolet-Visible Spectrophotometry (UV-Vis) and X-Ray Diffraction (XRD). Finally, time-kill curve study of Candida glabrata and H+-ATPase activity of this yeast were evaluated in the presence of biosynthesized silver NPs. The data obtained from SEM shows that Candida albicans can synthesize silver NPs with the size of 20 to 80 nm with spherical to oval morphology and XRD showed that these NPs are crystalline in nature. Also, the data obtained from the Time-kill curve study of Candida glabrata and the study of H+- ATPase activity of this yeast indicated that the cells of this yeast were reduced at (P ≤ 0.001) level and indicated favourable antifungal activity on this yeast. In fact, it can be concluded that Candida albicans is potentially able to synthesize silver NPs and these NPs have favourable antifungal properties. In addition, it was revealed that standard strain of Candida glabrata is more sensitive than clinical strain to biosynthesizes silver NPs.
Keywords
Biosynthesis of NPs, Silver NPs, Antifungal effects, Candida albicans, Candida glabrata
Introduction
Controlling the growth of microorganisms is one of the important issues paid attention to by World Health Organization (WHO) and the related authorities in every country and these authorities seek to employ the best and most effective method for controlling these pathogens [1]. Regardless of the herbal medicines that are less effective than before due to the resistance of microorganisms, currently mot substances that are used for treatment of the infections caused by microorganisms have chemical nature that have side effects [2-4]. On the other hand, microbial resistance is a serious and global problem that has challenged the treatment of infections with antibiotics [5,6]. Emergence of the science of nanotechnology in 21st century has opened a new window for the researchers. One of the main products of this technology is metal particles in the range of 1 to 100 nm that are referred to as metal nanoparticles. The unique physical, chemical, electrical, magnetic and catalytic properties of nanoparticles that are dependent on their size have resulted in their application in different fields [7,8]. Recently, researchers in the field of health have obtained favourable results by applying these substances against infectious agents. One of these nanoparticles is silver NPs. Due to their unique properties, silver NPs are considered an appropriate candidate for preventing and controlling the pathogenic microorganisms [9-11]. Different methods have been proposed for synthesis of silver NPs including chemical reduction, sonochemical synthesis, solochemical synthesis, mechanochemical synthesis and hydrothermal method [12-15]. Using these methods for synthesis of silver NPs requires chemical reactants and high temperature cycle, in addition to being time consuming. As the reactants used for the synthesis and control of the size of these NPs have chemical nature in most of these methods, these methods results in pollution both for humans and the environment [16,17]. Researchers’ efforts for solving such problems have resulted in development of new methods based on biological living systems for synthesis of NPs in recent years. In other words, with the aid of living systems such as microorganisms and plants, researchers have been able to synthesis nanomaterials that, in addition to having favourable medical properties, have no toxicity for human body and are completely compatible with the environment. The synthesis of NPs with the help of microorganisms has been proved by many researchers [18,19]. On the other hand, yeast infections including candidiasis are among the most common fungal infections that are created by different Candida species. Candidiasis is not just a diseases; it is referred to a wide range of diseases that are caused by different Candida species. Among the identified Candida species, Candida albicans is the cause of 60 per cent of Candida infections. However, other Candida species such as Candida glabrata, due to being resistant to azole medications, have widely increased and result in death in most of the cases. The inherent resistant against antifungal medications that exists in this microorganism has created challenge for the control and treatment of symptomatic infections caused by this yeast [20-22]. Considering the resistance of these organisms against azole antifungal medications and advancing nanotechnology, a method can be provided to treat these infections with using themselves. In this study, silver NPs were synthesized from Candida albicans and their in vitro antifungal properties against yeast cells of as Candida glabrata were assessed.
Materials and Methods
The materials and mediums used in this study such as silver nitrate, Sabouraud Dextrose Agar (SDA), Sabouraud Dextrose Broth (SDB), CHROM agar and Corn Meal Agar were purchased from Merck Company of Germany. The standard strain of Candida glabrata (ATCC: 90030) was purchased from Center of Scientific and Industrial Research of Iran. The silver nanoparticles in this study that were synthesized from the clinical strain of Candida albicans separated from patients with weak immune system, that were proved to be able to reduce silver ion and had potentiality to reduce Ag+ to Ag0, and their antifungal activity were assessed on the cells of the standard and clinical strains of Candida glabrata [23].
Biosynthesis of silver nanoparticles
First, the clinical strain of Candida albicans was cultured on SDA medium at 37°C for 24 hours. After 24 h of incubation, the grown colonies were cultured in an Erlenmeyer flask containing 29 ml of SDB, in order to obtain cell mass, and were kept in shaking incubator with 200 rpm speed and at 37°C. Then, using a centrifuge with the speed of 6000 rpm, the cells were separated from the medium and were washed with Phosphate buffered saline with pH=7.4. Then, 2 g of the wet weight of the yeast was removed and added to 5 ml of 1.5 mM silver nitrate and transferred to a sterile tube. The tubes were kept in darkness and at room temperature. After 2 h, and after witnessing the change in color and ensuring the transformation from Ag+ to Ag0, the tube containing cell biomass and silver nitrate solution was kept in shaking incubator again for 48 h to separate cell biomass. Then, yeast biomass was separated using a centrifuge with the speed of 6000 rpm and was used for SEM, UV-vis and XRD tests in the subsequent stages [23].
The study of the physicochemical characteristics of the biosynthesized nanoparticles
Scanning electron microscope (SEM): SEM was used for determining the shape and the size of the biosynthesized silver nanoparticles. In order to prepare the necessary sample, first, 2 g of the wet weight of the separated biomass together with Glutaraldehyde 2.5% were incubated at temperature of 4°C for 4 h. It was placed in room temperature with Osmium tetroxide 1%, after the incubation. Dehydration was done with alcohol after the solidification of the sample and then resin solution and oxypyrroline were added and the sample was kept at 60°C for 24 h. Finally, the prepared sample was assessed under SEM [23].
UV-Visible spectroscopy: The absorption spectrum of the biosynthesized nanoparticles was explored in UV-Vis spectrophotometer. For this purpose, 50 μl of the separated solution was added to the cuvette by 50 μl of distilled water and this spectroscopy was done in UV-Vis device at the wavelength of 300 to 800 nm [24].
XRD analysis: After the completion of biosynthesis of nanoparticles by Candida albicans, the resulted colloidal solution was centrifuged with 18000 rpm speed for 15 minutes. Afterwards, the supernatant was removed. For dispersing the precipitated nanoparticles, washing with double distilled water was done and centrifugation was repeated three times. And after each time of centrifugation, the precipitated nanoparticles were washed with double distilled water. Finally, the obtained suspension was put on silicon wafer and, after being dried, was put under XRD analysis [24].
The study of antifungal properties of biosynthesized nanoparticles
The effect of the nanoparticles on the clinical strain and standard strain (ATTC: 90030) cells of Candida glabrata was assessed. The clinical isolates in this study were separated and identified using classical culture methods on Corn Meal Agar, Chrome Agar, biochemical tests of sugar absorption and germ tube formation from patients with weak immune system hospitalized in hospitals in Shiraz. Then, a suspension equivalent to 1 × 106 cells in each ml was prepared in SDB from each strain using standard methods and spectrophotometer device at wavelength of 530 nm.
Based on the results obtained from previous studies the concentration 0.37 × 108 particles ml-1 has had a favourable effect on Candida albicans [25]. Different concentrations 0.18 × 108 Particles ml-1, 0.37 × 108 Particles ml-1 and 0.74 × 108 Particles ml-1 were used in this study for creating a link between the consumed dose of silver nanoparticles and the antifungal effect. This concentrations 1 were calculated and prepared in deionized water using equation n1v1=n2v2. The prepared yeast and nanoparticle suspensions were kept in refrigerator for the next stages.
Time-kill curve study of Candida glabrata
Exploring this factor was done in the presence of 0.18 × 108 Particles ml-1, 0.37 × 108 Particles ml-1 and 0.74 × 108 Particles ml-1 concentrations of the biosynthesized nanoparticles on standard and clinical strains in 0-48 h time range. First, the yeast cells with the dilution of 1 × 106 cell ml-1 were mixed with the intended concentrations that were prepared in the previous stage. Then, the 100 μl was removed from the obtained suspension and cultured on sterile medium SDA. The plates were placed in incubator for 48 h at 37°C. During this time the grown colonies were counted based on Colony-Forming Unit (CFU) at 0, 2, 4, 6, 8, 10, 12, 24 and 48 h. The yeast suspension of Candida glabrata which was free from nanoparticles was used as control group [26].
Proton efflux measurement
The activity and measurement of proton pump in Candida glabrata cells of the standard strain and clinical isolate was done through measurement of (intracellular) pH of the external environment. For this purpose, the cells of the middle phase were removed from SDB and were washed with distilled water twice. Then 0.1 g of the yeast cells was added to 5 ml of the solution containing (0.1 M KCl, 0.1 mM CaCl2) and distilled water. The prepared suspension was kept in double-jacketed glass container with constant stirring. The initial pH was adjusted on 7 using 0.1 M HCl/NaOH. Afterwards, for exploring the impact of the biosynthesized nanoparticles on the activity of the proton pump, 100 μl of the nanoparticles with concentrations of 0.18 × 108 Particles ml-1 , 0.37 × 108 Particles ml-1 and 0.74 × 108 Particles ml-1 were separately added to the prepared suspension. Finally, the H+ extrusion rate was measured by the consumption of 0.01 N NaOH [27].
Intracellular pH (pHi) measurement
In order to measure intracellular pH, 0.1 g of Candida glabrata yeast cells (ATCC: 90030) was separated from SDB. These cells, after being washed in 5 ml of solution containing (0.1 M KCl, 0.1 mM CaCl2), were suspended and the intended volume as adjusted using distilled water. In order to explore the impact of biosynthesized silver nanoparticles on intracellular pH level, 0.18 × 108 Particles ml-1, 0.37 × 108 Particles ml-1 and 0.74 × 108 Particles ml-1 concentrations of these nanoparticles were used. The initial pH was adjusted on 7. The prepared suspension was kept in a shaking incubator with the speed of 200 rpm at 37°C for 30 min. The pH changes in suspension were measured using pH meter [27].
Statistical analysis
The results of this study were the outcome of three replicates with standard deviation error. These results were analysed using statistical software SPSS (version 19) and one way ANOVA was used for determining the significance of the tests.
Results
The physicochemical characteristics of the biosynthesized nanoparticles
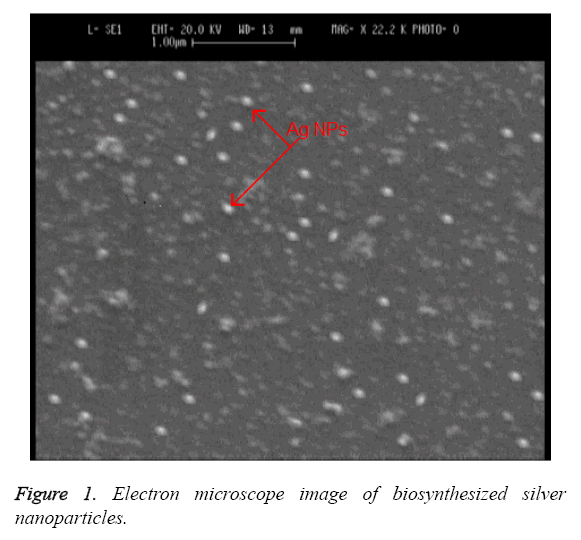
Electron microscope imaging: The observed change of color from milky white to red indicates the synthesis of silver nanoparticles by Candida albicans. A Scanning Electron Microscope (SEM) is used for exploring the size and the shape of these nanoparticles. Figure 1 shows the image of biosynthesized silver nanoparticles under SEM. The data obtained from this imaging indicates that the nanoparticles are synthesized from Candida albicans in sizes of 20 to 80 nm with spherical and oval morphology and in extracellular form.
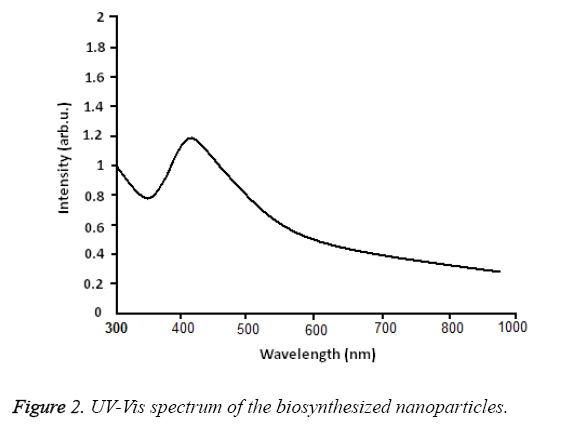
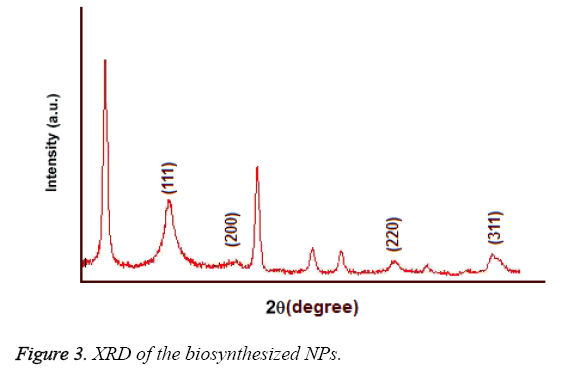
UV-Vis spectroscopy: Figure 2 shows the UV-Vis spectroscopy of the biosynthesized nanoparticles at the wavelength of 300 to 800 nm. The surface Plasmon resonance of the nanoparticles in the range of 400 to 450 nm has resulted in the creation of an absorption peak in 420 nm area which indicates the existence of silver nanoparticles in this area. Figure 3 shows the X-ray diffraction for the nanoparticles biosynthesized by Candida albicans. Increase in the sharpness of the XRD peaks in the Miller indices of the surfaces (111), (200), (220) and (311) that are related to angles 38.43, 46.255, 64.51 and 70.011 respectively indicates that the synthesized nanoparticles have a crystalline nature.
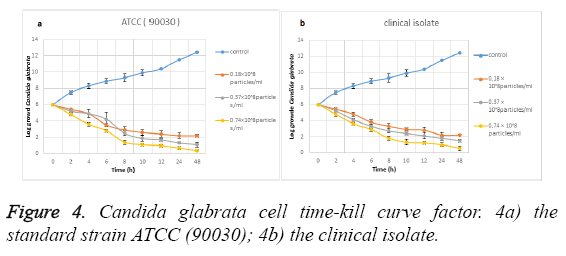
Time-kill curve study: Figure 4 shows the effect of the biosynthesized silver NPs on Candida glabrata time-kill curve. Figure 4a shows the effect of these NPs on the standard strain (ATCC: 90030) and Figure 4b shows the effect on clinical isolate. The data analysis shows that the silver NPs synthesized from Candida albicans have a favourable effect on Candida glabrata cells at the significance level of (P ≤ 0.001).
In a way that its antifungal activity is increased with the increase of the consumed dose. In fact, among the explored concentrations the most effective concentration is related to 0.74 × 108 particles ml-1. The analysis of the data of this study indicates that the Candida glabrata clinical strain cells showed a significant decrease in the presence of the nanoparticles in the aforementioned concentration in a way that it is reduced from Log 106 to Log 102. The effect on Candida glabrata standard strain (ATCC: 90030) cells is more in a way that these cells have been reduced from Log 106 to Log 10. Therefore the results indicate that the standard strain of Candida glabrata is more sensitive than the clinical strain, in the presence of the synthesized nanoparticles. In fact, this can be interpreted to indicate that the biosynthesized NPs, due to having crystalline aspects especially at levels (111) and (200), have a high electron density that has resulted in the bonding between these nanoparticles and the yeast cells.
Proton efflux measurement: Proton pump is an event dependent on H+ ATPase in fungal cells that is involved in pH adjustment and cell growth adjustment, by consuming energy, especially in food stress condition. Also, intracellular pH adjustment by proton pump has a critical role in the level of natural growth of Candida cells. The activity of proton pump and pH changes was explored at the presence of biosynthesized silver nanoparticles in this study. Table 1 shows the data related to the measurement of Proton efflux pump. Data analysis showed that the levels of proton pump retention in the concentrations of 0.18 × 108 Particles ml-1, 0.37 × 108 Particles ml-1 are equal to 39 and 56 per cent respectively in the standard strain (ATCC: 90030) (P ≤ 0.001) and equal to 36 and 53 per cent respectively in the clinical isolate (P ≤ 0.001). The concentration of 0.74 × 108 Particles ml-1 results in the level of retention of Proton pump being equal to 69 and 65 per cent in the standard and clinical strains respectively at level (P ≤ 0.001). This effect indicates the antifungal effect of the nanoparticles is increased with the increase of the used dosed. Also, the data in Table 1 shows that these nanoparticles have the highest level of impact on the standard strain (ATCC: 90030) of Candida glabrata, compared with the clinical isolate.
| Treatment | Range of relative H+ efflux rate (× 10-11 moles min-1 mg cells) | |
|---|---|---|
| ATCC (90030) | Clinical | |
| Control | 1 | 1 |
| 0.18 × 108 Particles ml-1 | 1.4 ± 0.2 (39%) | 1.29 ± 0.1 (36%) |
| 0.37 × 108 Particles ml-1 | 2.06 ± 0.5 (56%) | 1.94 ± 0.2 (53%) |
| 0.74 × 108 Particles ml-1 | 2.5 ± 0.3 (69%) | 2.42 ± 0.1 (65%) |
Table 1: The effect of biosynthesized silver nanoparticles on the activity of Candida glabrata proton pump.
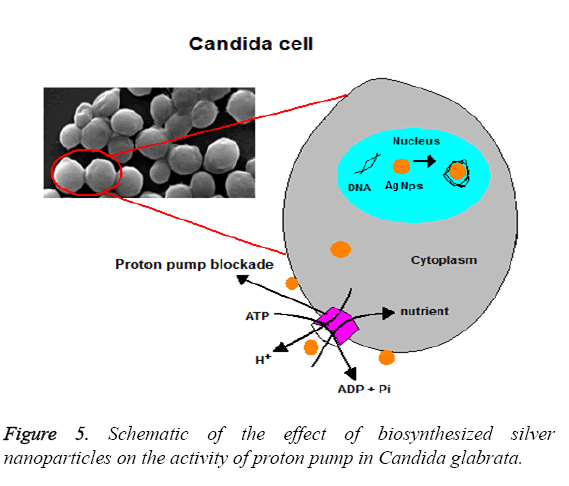
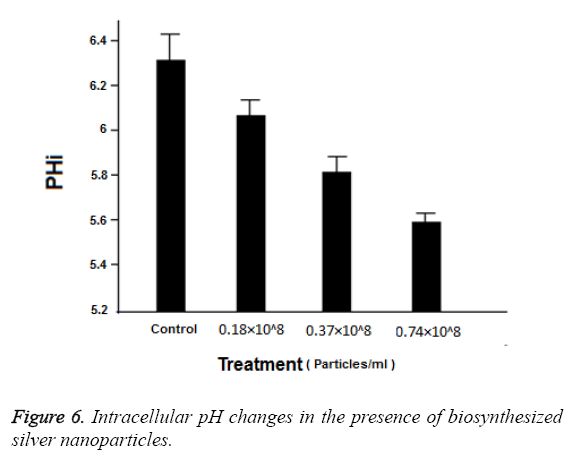
Figure 5 shows the effect in a schematic form. Normally, intracellular pH is adjusted between 6 and 7 by the activity of H+ ATPase. Figure 6 shows the changes of intracellular pH in the presence of silver nanoparticles. Also, it is evident in this Figure that the intracellular pH in the control group is equal to 6.3 and the highest level of effects related to 0.74 × 108 Particles ml-1 which is equal to 5.6 (P ≤ 0.001). While the concentrations 0.37 × 108 Particles ml-1 and 0.18 × 108 Particles ml-1 had less effect at the levels (p ≤ 0.001) and (p=0.001) respectively. These results indicate the favourable antifungal effect of biosynthesized silver nanoparticles.
Discussion and Conclusion
The resistance of the microbes and its emergence as a global problems resulted in efforts for findings an effective drug compound against the microorganisms. In line with this, many compounds were used against microorganisms by researchers. Silver compounds are among materials that their antibacterial effects were proven by researchers more than other compounds [28]. Also, the combination of silver with other materials resulted in the increase of the antimicrobial effect of the new compound. For example, the combination of silver nitrate with sulfadiazine and production of silver sulfadiazine cream resulted in the increase of the effect of sulfadiazine against Staphylococcus aureus and Escherichia coli [29]. Gradually, researchers found that with the increase of surface to volume ratio the effect changed and the antimicrobial effect was increased. This resulted in the emergence of particles called nanoparticles with a dimension of 1 to 100 nm [30].
With the application of silver nanoparticles as an effective drug compound against microorganisms, different methods were invented for the synthesis of these nanoparticles in a way that the invention made it possible for researchers to synthesize nanoparticles with a desired morphology and size and with favourable effect on microorganisms. Chemical and physical synthesis methods are the main and most common methods for production of silver nanoparticles. However, chemical methods for synthesis of silver nanoparticles result in toxic effects on human cells and pollution in water, soil and air due to the nature of the chemical reactants and stabilizers in synthesis stages. On the other hand, as these materials enter the food chain in living ecosystems, irreparable damages will occur. And physical methods have limitations as there is a need for a high temperature cycle (1000°C) and there are complexities in synthesis stages. In addition, the risk of potential explosion and fire of metal nano powder in the air makes the researchers search for alternatives to the present methods for synthesis of nanoparticles.
The challenges in this regard made researchers search for simple and more cost-effective ways for synthesis of silver nanoparticles that, in addition to having favourable effects on infectious agents, are less toxic to the body and the environment. The researchers found that methods for biosynthesis of nanoparticles can have such characteristics. The invention of the biosynthesis method resulted in a new era for studies all over the world on this field. In this regard, the study by Sadhasivam et al., can be pointed out. They synthesized silver nanoparticles from Streptomyces hygroscopicus [31]. In addition to the biosynthesis of silver nanoparticle by bacteria, fungi are potentially able to reduce silver ions and produce nanoparticles. In this regard, Verma et al., biosynthesized silver nanoparticles with dimensions of 10 to 20 nm from Aspergillus clavatus [32]. One of the problems in the biosynthesis of nanoparticles by fungi is the higher amount of time needed due to the slower growth of the fungi compared with bacteria. In their study titled the synthesis of gold nanoparticles from Fusarium oxysporum, Mokerji et al., pointed out that the biosynthesis of nanoparticles by fungi needs 120 h while bacteria can synthesize nanoparticles in a few hours [33]. In another study, Kathiresan et al., synthesized silver nanoparticles in dimensions of 1 to 100 nm from Penicillium fellutanum [34].
In comparison with the conducted studies, the present study used Candida albicans for synthesis of silver nanoparticles as it grows in a fast and simple way. The data in this study indicated that the yeast can synthesize silver nanoparticles with dimensions of 20 to 80 nm with crystalline nature. Preparation of cell biomass in less than 24 h and the change of color observed 2 h after the beginning of the reaction indicates that the biosynthesis of silver nanoparticles is done faster by this yeast, compared to other fungi. The costs o f obtaining nanoparticles is one of the limitations of chemical synthesis of nanoparticles and this limitation is highly reduced using biosynthesis methods because in most methods for nanoparticle biosynthesis the nanoparticles are synthesized in an extracellular way and there is no need for unnecessary compounds for obtaining the nanoparticles and these nanoparticles can be directly used for treatment purposes [35]. In the present study were synthesized from Candida albicans in an extracellular form and were directly used against the pathogenic yeast Candida glabrata.
Data analysis indicated that these nanoparticles have a favourable antifungal effect on the aforementioned yeast in a way that they resulted in a significant reduction of the yeast cells in 0 to 48 h at P ≤ 0.05 levels. The effect of silver nanoparticles against different Candida species has been verified by other researchers. In one study, Nozari et al., used silver nanoparticles together with Fluconazole against Candida albicans, Candida glabrata, Candida krusei and Candida tropicalis. Their results indicated that silver nanoparticles result in the antibiotic effect of Fluconazole [36]. In comparison with the aforementioned study, we synthesized silver nanoparticles from Candida albicans and used them against Candida glabrata. The data analysis indicated the antifungal effect of these nanoparticles. The difference between the sensitivity of the standard and clinical strains of Candida glabrata against the biosynthesized silver nanoparticles in this study indicates that the antifungal properties of nanoparticles is dependent on the strain type, in addition to dose and time, in a way that it was revealed that the clinical strain of Candida glabrata, compared with the standard strain, is more resistant to nanoparticles.
In this study it was tried to produce silver nanoparticles in an extracellular way, with the aim of creating an optimal condition compared with chemical and physical methods, and to assess their antifungal properties. Based on the results of the study it can be concluded that Candida albicans is potentially able to synthesize nanoparticles and due to the growth speed and ease of culture, it is an appropriate option, compared with other microorganisms, for synthesis of silver nanoparticles. These nanoparticles have favourable antifungal properties. It is recommended that researchers assess these results on animal models too for more clarification of the antifungal activity of nanoparticles. Because produced nanoparticles have a high ratio of surface area to volume, they exhibit special surface properties that are different from other bulk peers materials, So far a thorough understanding has not been achieved for harmful effects of used nanoparticles in clinical studies; This large surface area of nanoparticles allow more contact with the cell membrane, and also gives them a greater capacity to attract and immigration. This means that intensity and toxicity profiles of nanoparticles cannot be extrapolated from their bulk state information. In addition, nanosilvers showed toxic effects on many multicellular organisms such as mice and human. Some evidences suggests that nanosilvers after entering into the body, can accumulate in the body or lead to damage in some tissues such as liver, lungs and even into the bloodstream and crossing from Blood-Brain Barrier (BBB); And if nanosilvers can transform to bulk silver, serious and widespread risks of these substances is cleared. However, more comprehensive studies are required to evaluate the effects of silver nanoparticles in clinical activities.
References
- WHO. How to investigate drug use in health facilities (selected drug use indicators). WHO/DAP/93.1 Geneva WHO, 1993.
- Lemar KM, Passa O, Anon MA, Cortassa S, Muller CT, Plummer S. Allyl alcohol and garlic (Allium sativum) extract produce oxidative stress in Candida albicans. Microbiol 2005; 151: 3257-3265.
- Wiland P, Szechcinski J. Proximal tubule damage in patients treated with gentamicin or amikacin. Pol J Pharmacol 2003; 55: 631-637.
- Marambio-Jones C, Hoek E. A review of the antibacterial effects of silver nanomaterial’s and potential implications for human health and the environment. J Nanopart Res 2010; 12: 1531-1551.
- Aguiar JM, Chacon J, Canton R, Baquero F. The emergence of highly fluoroquinolone-resistant Escherichia coli in community-acquired urinary tract infections. J Antimicrob Chemother 1992; 29: 349-350.
- Goldmann DA, Weinstein RA, Wenzel RP, Tablan OC, Duma RJ. Strategies to prevent and control the emergence and spread of antimicrobial-resistant microorganisms in hospitals. Jama 1996; 275: 234-240.
- Wijnhoven S, Peijneburg W, Herberts C. Nano-silver-a review of available data and knowledge gaps in human and environmental risk assessment. 2009; 3: 109-138.
- Saber MH, John JS. Toxicological highlight safety evaluation of silver nanoparticles. Toxicological Sci 2009; 108: 223-224.
- Hussain SM, Schlager JJ. Safety evaluation of silver nanoparticles: inhalation model for chronic exposure. Toxicol Sci 2009; 108: 223-224.
- Li WR, Xie XB, Shi QS, Zeng HY, Ou-Yang YS. Antibacterial activity and mechanism of silver nanoparticles on Escherichia coli. Appl Microbiol Biotechnol 2010; 85: 1115-1122.
- Yu B, Leung KM, Guo Q, Lau WM, Yang J. Synthesis of Ag-TiO2 composite nano thin film for antimicrobial application. Nanotechno 2011; 22: 115603.
- Shrivastava S, Jyung WO, lungue M. Characterization of enhanced antibacterial effects of nano silver nano particles. J Nanotechnol 2010; 25: 103-120.
- Esmaielzadeh KA, Farzalipour TM, Pourabbas B. Sonochemical Synthesis of Zno Nanoparticles. E Temp Sonic Pow Mat Res Bul 2008; 43: 648-654.
- Sankapal BR. Chemical and electrochemical synthesis of nanosized tio2 anatase for large-area photon conversion. CR Chimie 2006; 9: 702-707.
- Gajovic A. Mechanochemical preparation of nanocrystalline TiO2 powders and their behavior at high temperature. J All Comp 2005; 398: 188-199.
- Vaezi MR, Sadrnezhad SK. Nanopowder Synthesis of Zinc Oxide via Solochemical Processing. J Mat Des 2007; 28: 515-519.
- Barbu E, Molnar E, Tsibouklis J, Gorecki DC. The potential for nanoparticle-based drug delivery to the brain: overcoming the blood-brain barrier. Expert Opin Drug Deliv 2009; 6: 553-565.
- Bystrzejewska PG, Golimowski J, Urban PL. Nanoparticles-Their potential toxicity, waste and environmental management. Waste Manag 2009; 29: 2587-2595.
- Forough M, Farhadi KH. Biological and green synthesis of silver nanoparticles. Turkish J Eng Env Sci 2010; 34: 281-287.
- Husseiny MI, El-Aziz MA, Badr Y, Mahmoud MA. Biosynthesis of gold nanoparticles using Pseudomonas aeruginosa. Spectrochim Acta A Mol Biomol Spectrosc 2007; 67: 1003-1006.
- Boerlin P, Boerlin PF, Ddurussel C. Cluster of atypical Candida isolates in a group of human immunodeficiency virus positive drug users. J clin microbiol 2002; 40: 2614-2621.
- Fidel PL, Vazquez JA, Sobel JD. Candida glabrata: review of epidemiology, pathogenesis, and clinical disease with comparison to C. albicans. J Clin microbiol rev 1999; 12: 3-69.
- Wingard JR. Importance of Candida species other than C. albicans as pathogens in oncology patients. Clin Infect Dis 1995; 20: 115-125.
- Rahimi GH, Alizade F, Khodavandi A. Mycosynthesis of silver nanoparticles by Candida albicans and evaluation antibacterial activity on Staphylococcus aureus and Escherichia coli . Trop J pharma sci 2016; 2: 764-773.
- Kalimuthu K, Suresh Babu R, Venkataraman D, Bilal M, Gurunathan S. Biosynthesis of silver nanocrystals by Bacillus licheniformis. Colloids Surf B Biointerf 2008; 65: 150-153.
- Masoud N, Shahin O, Ali EZ, Seyed AM, Gholamreza M, Yazdan M, Ghasem R. Plant synthesis of silver nanoparticles using Matricaria chamomilla plant and evaluation of its antibacterial and antifungal effects. J Biomedical Research 2014; 26: 794-799.
- Rahimi Gh, Khodavandi A, Jannesar R, Alizadeh F, Yaghobi R, Sadri A. Evaluation of antifungal effects of ethanolic and aqueousextracts of Zataria multiflora herb in the pathogenic yeast Candida albicans biofilm inhibition. J pure appl microbiol 2014; 8: 4559-4564.
- Wani IA, Ahmad T, Manzoor N. Size and shape dependant antifungal activity of gold nanoparticles: a case study of Candida. Colloids Surf B Biointerf 2013; 101: 162-170.
- Moyer CA, Brentano L, Gravens DL, Margraf HW, Monafo WW. Treatment of large human burns with 0.5% silver nitrate solution. Arch Surge 1965; 90: 812-825. Stanford W, Rappole BW, Fox CL. Clinical experience with silver sulfadiazine, a new topical agent for control of pseudomonas infections in burns. J Trauma 1969; 9: 377-388.
- Guzman M, Dille J, Godet S. Synthesis and antibacterial activity of silver nanoparticles against gram-positive and gram-negative bacteria. Nano medicine Nanotechnol Biol Med 2012; 8: 37-45.
- Sadhasivam S, Shanmugam P, Yun K. Biosynthesis of silver nanoparticles by Streptomyces hygroscopicus and antimicrobial activity against medically important pathogenic microorganisms. Colloids Surfaces B Biointerfaces 2010; 81: 358-362.
- Verma VC, Kharwar RN, Gange AC. Biosynthesis of antimicrobial silver nanoparticles by the endophytic fungus Aspergillus clavatus. Nanomedicine (Lond) 2010; 5: 33-40.
- Mukherjee P, Senapati S, Mandal D, Ahmad A, Khan MI. Extracellular synthesis of gold nanoparticles by the fungus Fusarium oxysporum. Chembiochem 2002; 3: 461-463.
- Kathiresan K, Manivannan S, Nabeel MA, Dhivya B. Studies on Silver Nanoparticles Synthesized by A marine fungus, Penicillium fellutanum isolated from coastal mangrove sediment. Colloids Surf B 2009; 71:133-137.
- Murali S, Absar A, Islam Khan M, Kumar R. Biosynthesis of metal nanoparticles using fungi and Actinomycete. Curr Dir Psychol Sci 2003; 1201-1207.
- Nozari SH, Haydari kohan F, Ashrafi khozani M, Ahmadi F, Ghasemi Z, Nami N, Falahati F. Comparision of antifungal effect of fluconazole alone and incombination with nanosilver particles against candida species isolated from chronic candida vulvo vaginitis. Razi J Med Sci 2012; 18: 9-14.