ISSN: 0970-938X (Print) | 0976-1683 (Electronic)
Biomedical Research
An International Journal of Medical Sciences
Research Article - Biomedical Research (2017) Volume 28, Issue 17
Recognition of finger movements using EEG signals for control of upper limb prosthesis using logistic regression
Amna Javed, Mohsin I. Tiwana*, Moazzam I. Tiwana, Nasir Rashid, Javaid Iqbal and Umar Shahbaz Khan
Department of Mechatronics Engineering, National University of Sciences and Technology, H-12, Islamabad, Pakistan
- *Corresponding Author:
- Mohsin I. Tiwana
Department of Mechatronics Engineering
National University of Sciences and Technology
Islamabad, Pakistan
Accepted date: June 02, 2017
Brain computer interface decodes signals that the human brain generates and uses them to control external devices. The signals that are acquired are classified into movements on the basis of feature vector after being extracted from raw signals. This paper presents a novel method of classification of four finger movements (thumb movement, index finger movement, middle and index finger combined movement and fist movement) of the right hand on the basis of EEG (Electroencephalogram) data of the movements. The data-set was obtained from a right-handed neurologically intact volunteer using a noninvasive BCI (Brain-Computer Interface) system. The signals were obtained using a 14 channel electrode headset. The EEG signals that are obtained are first filtered to retain alpha and beta band (8-30 Hz) as they contain the maximum information of movement. Power Spectral Density (PSD) is used for analysis of the filtered EEG data. Classification of the features is done using various classifiers. Various classifiers have been tested and compared on basis of the mean class accuracy achieved. The classifier chosen for the study is logistic regression, which gives an accuracy of 65%. Other classifiers that were tested were multi-layer perceptron, linear discriminant analysis, and quadratic discriminant analysis. The novelty of this research is the targeted finger movements. These movements were targeted so they can be further used for control of upper limb prosthesis.
Keywords
Brain computer interface (BCI), Logistic regression, Power spectral density (PSD), Electroencephalogram (EEG), Mu and beta rhythms
Introduction
Brain Computer Interface (BCI) uses signal of brain function to enable individuals to communicate with the external world, bypassing normal neuromuscular pathways [1]. The main purpose of developing such a system is to provide those people with a method of communication with the external world, whose normal muscular pathways have been permanently damaged by disorder such as ALS (Amyotrophic Lateral Sclerosis) or an injury to the spinal cord. These brain signals are called Electroencephalographic (EEG) signals and after some pre-processing, they can be used to detect the user’s intent for a particular movement of any part of the body.
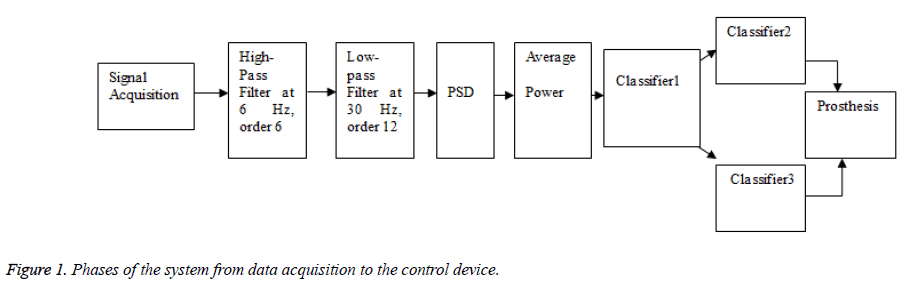
BCI system is like any communication system and has an input, i.e. the EEG of the user, an output (i.e. device commands), and components that translate input into output. The EEG is acquired via a headset that consists of electrodes to capture the signals. These signals are transformed into the digital domain and sent to a computer for signal processing. The signal processing consists of digital filtration, feature extraction and a translating algorithm (classifier). After the intent of the person has been identified by the translating algorithm, the signal is sent via a control interface to the control device [1]. The phases of the system that was used in this research are shown in Figure 1. It is important that any feature chosen should have a large signal to noise ratio. There are a variety of feature extraction procedures such as spatial filtering, voltage amplitude measurements, spectral analyses. These signals are then processed to show the user intent. In the end, the system is interfaced with a device for control.
This study uses power spectral density of finger movements of one hand occurring over the motor cortex as a feature to classify them [2]. The mu and beta rhythms that occur over the motor cortex have a frequency range of 8-30 Hz. These rhythms provide us with the information related to movement. Mu and beta rhythms are largest in amplitude when there is no movement and the amplitude decreases with movement (Event Related De-synchronization and Event Related Synchronization). These are useful for classification of different limb movements. Their amplitude can also be controlled with some training of the subject and hence, the system performance can increase [3]. PSD describes how the power of the signal is distributed over its frequency. There are many properties in PSD of brain signals that can be used as a feature. One of the most commonly used features is band power of a specific frequency band from the PSD of the signal. This study is using the average band power of the PSD of the mu and beta rhythms as a feature vector.
A classifier is applied to distinguish between different movements classes using the selected features. The classifier output is then transformed into an appropriate signal used to control a variety of devices. The training data helps the classifier build a model to differentiate between the classes. Some of the mostly used classifiers in BCI are Linear Discriminant Analysis, Neural Networks, Quadratic Discriminant Analysis and Support Vector Machines [4]. Some of these classifiers and the logistic regression model were trained and tested to classify the feature and their results were compared. ‘Matlab’ and ‘Weka’ were used for performing this task.
The movements of the fine body parts are not well-researched in EEG based BCI [5]. Researchers have concentrated on spatially apart parts such as arms, legs, tongue. Those who have classified finger movements of the same hand have used Electrocorticography (ECoG) signals that have a higher signal to noise ratio (SNR). However, those who have done similar work are mentioned in Table 1 along with the accuracy achieved in their study.
| Researchers | |||
|---|---|---|---|
| Name | Body part targeted | Electrode system | Accuracy |
| Xiao and Ding [6] | Ten fingers pairs | 128-Channel Geodesic EEG System 300 |
45.2% |
| Xiao and Ding [3] | Ten finger pairs | 128-Channel EEG sensor layout with 50 electrodes | 77% |
| Vuckovic and Sepulveda [7] | Right and left hand movement | 64-Channel biosemi active two systems. | 69% |
| Mohamed, Marwala and John [8] | Wrist and finger movement discrimination | GSN 128 | 65%-71% |
| Lehtonen, Jylanki, Kauhanen and Sams [13] | Visual cue based left and right index finger movement | 32-channel amplifier (Brain Products GmbH, Germany). | 80% |
| Wang and Wan [14] | Press with the index and little fingers the corresponding keys in a self-chosen order and timing ‘self-paced key typing’. | 28 electrode 10/20 system | 86% |
Table 1. Related research.
RanXiao et al. used a 128 channel electrode system to decode individual finger movements. They calculated the PSD of the EEG data and decomposed it using principal component analysis and then used support vector machine for classification of ten pairs of finger movements [6].
Vuckovic et al. classified the flexion and extension of left and right wrists and used a classifier based on Elman’s neural network and achieved an accuracy of 69%. They used amplitude of Gabor coefficients calculated over 125 ms and 2 Hz time-frequency windows as feature vectors. They proposed a 2-stage 4 class classifier model for classification and the same model is employed by this study [7].
Mohamed et al. classified wrist and finger movements of both right and left hand. They also classified both imaginary and real movements. The data was pre-processed using filtration and independent component analysis. For feature extraction, they calculated power spectrum and then used Bhattacharyya distance for feature selection. They tested two classifiers, one based on Mahalanobis Distance (MD). The other classifier they tested was Artificial Neural Network (ANN). The average accuracy achieved in this study was 65% and 71% for MD and ANN classifier respectively [8].
Wang et al. classified left or right finger movements. Their algorithm was based on common spatial subspace decomposition and support vector clustering [9].
Lehtonen et al. classified left versus right hand index finger movements. Their classifier was trained online. They removed linear trends from EEG data and then computed the Fast Fourier transform and filtered different frequency bands. They used temporal features for low frequencies (below 3 Hz) and instantaneous amplitudes for higher frequencies. The researchers used Kolmogorov-Smirnov (KS) test statistic for feature selection. Their classification was based on three linear transformations of the feature space and a linear classifier with a logistic output function [10].
Methodology
Experimental protocol and data acquisition
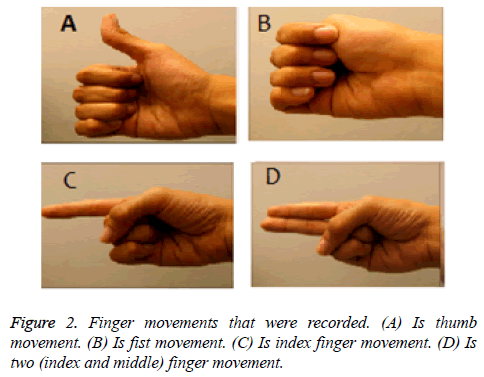
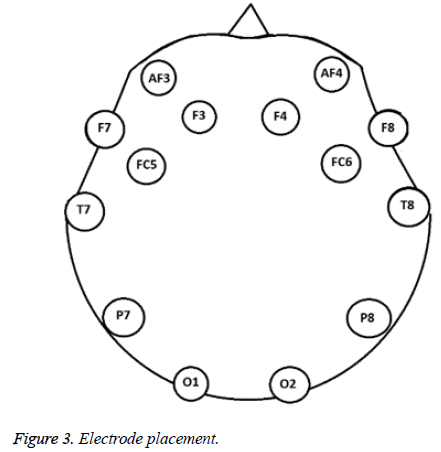
The subject was a healthy, right-handed, 22 y old female without prior training of the experimental procedure. The data acquisition was carried out in a closed room with the subject sitting comfortably in a chair with arms rested on the sides. The subject was asked to perform the movements shown on a computer screen that were displayed as a video. The video showed a person performing the movements that the user had to mimic. Each movement lasted 10 seconds and one trial was of 1 min 15 s with a 2 s cue between each movement. In the start of the video, there are 10 s of instructions for the volunteer. Then there is a 2 s cue in form of a star and then the hand movement to be performed is shown which the volunteer has to replicate. The movements that were recorded are shown in Figure 2. These movements were recorded via an Emotiv electrode headset that has 14 channels based on the international system which are AF3, F7, F3, FC5, T7, P7, O1, O2, P8, T8, FC6, F4, F8 and AF4 [2-4,9-16]. Signals were recorded at 128 Hz sampling rate (Figure 3).
The recording was done in 3 sessions. Each session had 20 trials with a break of a min between each trail and a 5 min break after 10 trails. One session took around 1 h. In the end, there were 60 recordings of each movement. 13 recordings of each movement had to be discarded because of improper electrode connection. During processing, 5 s of data of each movement is being used.
The data is truncated to 5 s and then it is filtered and processed further. The middle 5 s of data was taken, i.e. 2.5 s of data was truncated from both ends. This was done in order to ensure that only the true movement data was taken because when there is a shift from another movement, the volunteer doesn’t change movement instantaneously.
The volunteer is performing movements repetitively for the 5 s that are being processed. In a BCI paradigm, it is common to take length of recordings with repetitive movements since ERD occurs with start of each motion and ERS occurs at the relaxation of motion. Mohamed et al. are a few examples of researchers who used a similar experimental protocol [5,8,10,15].
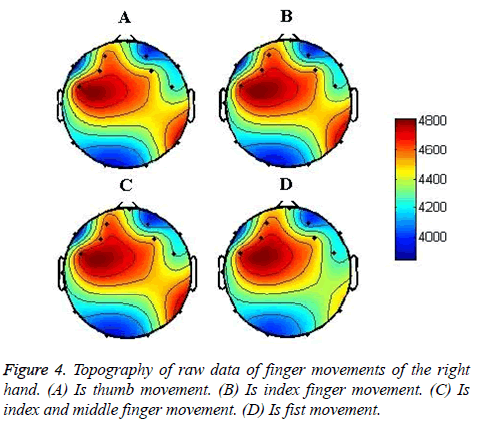
The EEG topography of raw data of each of the movements is shown in Figure 4. The plots are of a randomly taken data sample in which the channel values have been averaged over the 5 s window.
The average results are of the values as obtained from the CSV file produced by the Emotive headset. It should be noted that these values have a DC offset of around 4000 points. The units are in microvolts. The topography plots are interpolated by 2-D linear interpolation performed using “Mat lab”. It can be seen that all the movements are concentrated under the same region, which is the left hemisphere of the frontal lobe. This region is the sensorimotor cortex.
There are minor differences in the topography of the movements. This is because the locations of fingers on sensorimotor cortex lie close together and hence, are recorded by the same electrodes.
Even though emotive does not have electrodes in the region of major activity, interpolation of the channel data shows us the region of the brain where the activity is located.
Signal pre-processing
5 s of EEG data was digitally filtered using a Butterworth filter between 6-30 Hz. The high pass filter at 6 Hz had an order 6 and the low pass filter at 30 Hz had order 12. This was done because the alpha and beta band, which contains most movement related information, lies between 8-30 Hz. Butterworth filter gives a flat response with zero ripples and is used for this reason.
The eye artefact has a frequency of 2-5 Hz and is hence removed when a high pass filter of 6 Hz is applied. The reason of choosing alpha and beta signals for this study is that these signals are related to physical and mental relaxation. Theta and delta are related to unconscious, deep sleep or meditation [16].
Feature extraction
Power Spectral Density of the filtered signal was found by Welch’s method, which is a non-parametric method where PSD is calculated directly from the filtered signal itself. For each channel, the filtered signal is divided into 64 window segments with a 50% overlap. The 5 second data that was filtered is first divided into 64 windows. The window being used is a hamming window. Fast Fourier Transform is then applied to each windowed segment. The power spectral density of segment is then calculated.
The PSD of the each segment is given by equation 1 [12].

Where ‘Fs’ is the sampling frequency, ‘L’ is the length of segment and X (f) is the data after FFT has been applied on it. ‘U’ is the window normalization constant given by equation 2.

After the PSD of each segment is calculated, the corresponding values of each segment are added and averaged. The resultant vector is then averaged again and this gives us the average band power of the channel over 6-30 Hz band (as all other frequencies have been removed by filter). This process is repeated for each channel, which leaves us with a feature vector of 14 constituents showing the average band power of each channel.
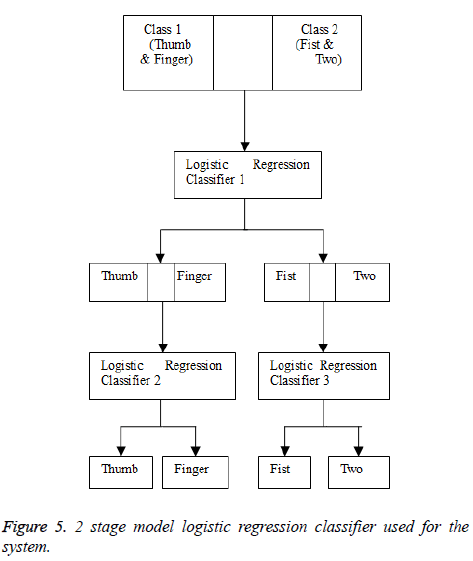
Classification method 1: 2 stage model
For classification, a 2 stage 4 class model was used (Figure 5) [17-19]. The classifier used is logistic regression. Logistic Regression is a type of probabilistic statistical classification model that estimates the probability of an event occurring. Unlike linear regression, it does not assume linear relationship between the dependant and independent variable.
In the first stage, the classification training was between two classes, one being the finger and thumb class and the other being the fist, index and middle finger (two) combined class. In the second stage, there are further two classifier networks. The first was trained to classify between finger and thumb class and the other was trained to classify between the fist and two (index and middle finger) class. ‘Weka 3.6.9’ was used to train the three models. The three models were trained by using 75% of the dataset. These trained models were then imported to ‘Matlab’ and were tested there. In “Matlab”, this study tested all the three stages with the remaining 25% of the dataset. “Weka” is data mining software that has a collection of machine learning algorithms and has tools for pre-processing as well as classification and more. In this proposed work, the suggested software for training of our classifiers is Weka. For the logistic regression classifier, the probability of the first class is given by Equation 3 [11].

Where F is the feature vector and B is the coefficients of logistic regression. Parameter F is the feature vector passed on by the feature extractor as described in section C.
Parameters BT are the coefficients of logistic regression calculated by Weka by the training of the classifier. Weka calculates the coefficients using Quasi-Newton method to find the optimized values. After the training stage, the classification equations are as under:
Initial classification: f=-1.4049+(feat(1) × -0.01539)+(feat(2) × 0.0333)+(feat(3) × 0.1447)+(feat(4) × -0.0591)+(feat(5) × -0.0644)+(feat(6) × 0.0589)+(feat(7) × -0.1048)+(feat(8) × 0.0672)+(feat(9) × -0.0073)+(feat(10) × -0.0466)+(feat(11) × -0.1013)+(feat(12) ×0.8546)+(feat(13) × 0.0012)+(feat(14) × -0.0025).
Thumb and finger classification: f1=0.6788+(feat(1) × -0.3851)+(feat(2) × 0.0593)+(feat(3) × 0.1621)+(feat(4) × -0.0054)+(feat(5) × 0.0642)+(feat(6) × -0.5581)+(feat(7) × 0.1561)+(feat(8) × 0.0018)+(feat(9) × -0.003)+(feat(10) × -0.0227)+(feat(11) × 0.1227)+(feat(12) × 0.1599)+(feat(13) × -0.0004)+(feat(14) × -0.0172).
Fist and two classifications: f1=-0.3847+(feat(1) × -0.0595)+ (feat(2) × 0.0193)+(feat(3) × 0.0361)+(feat(4) × -0.02)+ (feat(5) × -0.0458)+(feat(6) × 0.0377)+(feat(7) × -0.0856)+ (feat(8) × 0.0623)+(feat(9) × -0.0008)+(feat(10) × 0.008)+ (feat(11) × -0.1519)+(feat(12) × 0.9574)+(feat(13) × 0.0396)+ (feat(14) × -0.203).
The criterion for selection of class is: G(x)=class 1 if Pr>0.5, G(x)=class 2 if Pr<0.5. If Pr=0.5, the result is classified in neither classes and disregarded.
Other classifiers were also tested which are Multi-Layer Perceptron, Linear Discriminant Analysis (LDA) and Quadratic Discriminant Analysis (QDA). LDA and QDA were tested in a 2-stage 4 class model. For LDA and QDA, training was done for 75% of the dataset and they were tested over the remaining 25% dataset.
Discriminant analysis creates a hyper plane to separate the data into two classes. It assumes that different classes generate data based on different Gaussian distributions [14]. Both Quadratic (QDA) and Linear Discriminant Analysis (LDA) were trained in Matlab with a 75% training data set. Testing was done in Matlab with the remaining 25% data set. In LDA, it is assumed that the co-variance of each class is same and the mean varies while in QDA, both are assumed to vary [4]. The 75% and 25% training and testing division was done once and was used for logistic regression, LDA and QDA. In each division, the proportion of samples from each class was same for training and testing data.
The 2 stage model was also tested using 5 fold cross validation. In 5 cross validation, the data was divided into 5 disjoint subsets, each with equal proportions of samples of each class. 4 of these subsets are used for training and 1 for testing of data. This method is repeated 5 times with all possible combinations of 4 subsets used for training and 1 subset used for testing. The result is then averaged over 5 to give the end results.
Results
The confusion matrix is given in Table 2. Table 3 shows the accuracy of each classifier network in a 2-stage 4 class logistic regression classifier. Each class’s accuracy of classification by the entire classifier is shown in Table 4. The confusion matrix shows how many times the feature was correctly classified as its own class and how many times it was classified as any other class. For example, the thumb features were classified as thumb in 7 instances out of the total 13 instances tested, as finger in 1 instance, as fist in 3 instances and as two in 2 instances. Accuracies are calculated from the testing data set only. They were calculated from the confusion matrices shown by taking out the percentage of correctly classified instances for each class by total number of instances. For example, if out of 13 instances, thumb is correctly classified 7 times, the class accuracy is 7/13=55%. This is repeated for each class and the average of all is taken to give average per class accuracy, which in case of logistic regression is calculated as: (55+77+69+69)/400 × 100=67.5%. This is how accuracy is calculated for all classifiers. Table 5 shows the confusion matrix obtained from 5 fold cross validation with logistic regression in a 2-stage model. The mean class accuracy of the logistic regression 2-stage model is 67.5% with 75% training data set, while 5 fold cross validation test yields an accuracy of 44.27%.
| Class | Class the feature was classified as by the classifier | |||
|---|---|---|---|---|
| Thumb | Finger | Fist | Two | |
| Thumb | 7 | 1 | 3 | 2 |
| Finger | 3 | 10 | 0 | 0 |
| Fist | 0 | 0 | 9 | 4 |
| Two | 0 | 1 | 3 | 9 |
Table 2. Confusion matrix for 2 stage model logistic regression with 75% training 25% test set.
| Network number | Classification Accuracy with 75% training and 25% testing data. | Classification accuracy with 5 fold cross validation |
|---|---|---|
| 1 | 74% | 66% |
| 2 | 76% | 71% |
| 3 | 64% | 62% |
Table 3. Network classification accuracy of two-stage logistic classifier. Network (1) classifies between class 1(thumb + finger) and class 2(Fist + Two). Network (2) classifies between thumb and index finger. Network (3) classifies between two and fist.
| Class | Classification Accuracy with 75% training and 25% testing data | Classification accuracy with 5 fold cross validation |
|---|---|---|
| Thumb | 55% | 58% |
| Index finger | 77% | 39.5% |
| Fist | 69% | 52% |
| Index and middle finger | 69% | 27% |
Table 4. Per class accuracy using a 2-stage model logistic classifier.
| Class | Class the feature was classified as by the classifier | |||
|---|---|---|---|---|
| Thumb | Finger | Fist | Two | |
| Thumb | 28 | 4 | 9 | 7 |
| Finger | 9 | 19 | 6 | 14 |
| Fist | 4 | 4 | 25 | 15 |
| Two | 7 | 13 | 7 | 13 |
Table 5. Confusion matrix for 2-stage logistic regression with 5 folds cross validation testing.
Linear Discriminant Analysis classifier was also used as a 2- stage 4 class model and it was trained and tested in Matlab.
The mean class accuracy of this classifier was 31.25% with 75% training and 25% testing data set whereas 5 fold cross validation yields an accuracy of 24%.
Quadratic Discriminant Analysis was tested and trained as a classifier in Matlab. The classifier was used as the same 2- stage 4 class model.
The mean class accuracy of QDA was 31.47% with 75% training and 25% testing data set whereas mean class accuracy using 5 fold cross validation was 49%.
Table 6 compares the accuracy of each of the classifiers tested.
| Classifier | Mean Class Accuracy with 75% training and 25% testing data set. | Mean Class Accuracy with 5 fold cross validation |
|---|---|---|
| Logistic Regression | 67.5% | 44.27% |
| Linear Discriminant Analysis | 31.25% | 23.97% |
| Quadratic Discriminant Analysis | 31.47% | 48.9% |
Table 6. Comparison of classifiers for classification of finger movements with PSD as feature vector and 2 stage classification model used.
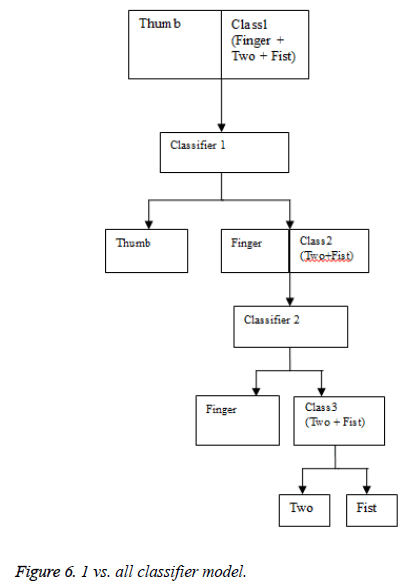
Classification method 2: 1 vs. all model
The second classification model that was tested was the 1 vs. all models. In this model, there are 3 classifiers in 3 stages.
Figure 6 shows the model flow chart. In the first stage, the classification is between the thumb and all the other classes. The next is between the finger and the remaining classes and in the final stage; the classification is between two and fist class. Logistic regression as well as QDA and LDA were tested in this manner.
As described previously, the classifier networks were tested by both 75% training and 25% testing split and 5 fold cross validation. In both cases, each division of data had the same proportion of samples from each class.
Logistic Regression was trained and tested in Weka 3.6.9 while LDA and QDA were trained and tested in Matlab.
Table 7 compares the classification accuracies of QDA, LDA and logistic regression in a 1 vs. all model with both 75% testing and 25% training method, and 5 fold cross validation method.
| Classifier | Mean class accuracy with 75% training and 25% testing data set | Mean class accuracy with 5 fold cross validation |
|---|---|---|
| Logistic Regression | 31% | 65% |
| Linear Discriminant Analysis | 36% | 43.36% |
| Quadratic Discriminant Analysis | 20% | 53.2% |
Table 7. Comparison of classifiers for classification of finger movements with PSD as feature vector and 1 vs. all model used.
Table 8 shows the confusion matrix of logistic regression as obtained from 5 fold cross validation method.
| Class | Class the feature was classified as by the classifier | |||
|---|---|---|---|---|
| Thumb | Finger | Fist | Two | |
| Thumb | 39 | 3 | 3 | 4 |
| Finger | 7 | 28 | 5 | 9 |
| Fist | 4 | 2 | 28 | 5 |
| Two | 1 | 7 | 9 | 32 |
Table 8. Confusion matrix for 1 vs. all logistic regression with 5 folds cross validation testing.
Table 9 shows the per class accuracy of 1 vs. all logistic regression model.
| Class | Classification Accuracy with 75% training and 25% testing data | Classification accuracy with 5 fold cross validation |
|---|---|---|
| Thumb | 25% | 80% |
| Index finger | 50% | 57% |
| Fist | 33% | 65% |
| Index and middle finger | 17% | 57% |
Table 9. Per class accuracy using a 1 vs. all model logistic classifier.
Discussion
From the results obtained from both the models and all the classifiers, it is seen that logistic regression in a 1 vs. all model gives the highest accuracy of 65% when tested with 5 fold cross validation. 5 cross validation is a much more reliable testing technique because in percentage split, it is possible that the split just happens to work better for a particular classifier. This was the case when the 2-stage logistic classifier was tested. Hence, to ensure reliability, each classifier was tested with both tests in the two models. The results prove logistic regression gives the highest accuracy results and should be used in the 1 vs. all model shown for the chosen movements. The confusion matrix of the classifier also shows that the classifier is not biased towards any class.
The research is using a low-cost EEG headset with only 14 channels with no electrode over the motor cortex. The accuracy for the 4-class problem, with movements that are closely located on the cortex, was not expected to be greater than 25% and accuracy greater than this show potential for work in this field. The impact of the work lies in the fact that an inexpensive headset can be used to classify a four-class problem of data from one hand. By using a headset with more electrodes that can capture the signals more accurately from the motor cortex, this methodology will show much better results and can be effectively used for prosthesis control.
Comparison with multi-layer perceptron
For further comparison, multi-layer perceptron was trained and tested by using 75% data set for training and remaining for testing and with 5 fold cross validation.
Multi-layer perceptron was trained and tested in “Weka”. This classifier is a feed forward artificial neural network classifier. It consists of an input layer, hidden layer and an output layer of computational nodes [13].
The multi-layer perceptron used had 9 hidden layers and each node had a sigmoid function. Using a multi-layer perceptron gives an accuracy of 44% with 75% training set. With 5 fold cross validation; MLP gives an accuracy of 54%.
Table 10 shows the per class accuracy of this classifier. It can be seen from the table that MLP is unable to classify the “fist” class.
| Class | Classification Accuracy with 75% training and 25% testing data | Classification accuracy with 5 fold cross validation |
|---|---|---|
| Thumb | 53% | 67% |
| Index Finger | 64% | 59% |
| Fist | 0 | 37% |
| Index and Middle Finger | 36% | 51% |
Table 10. Per class accuracy multi-layer perceptron.
Comparison with other works
Ran et al. classified 10 pair of finger movements and achieved an overall accuracy of 77%. If we compare pair accuracies with that achieved by RanXiao, the thumb vs. index finger was 71.27% while this study achieved 71% accuracy for that class. Our work is most closely related to RanXiao et al. research as they are concentrating on classification of individual finger movements.
The work of Vuckovic et al. basically proposed the usage of a 2-stage 4 class model for BCI systems and this study used their idea of a 2-stage model and applied it on our classifier.
Mohamed et al. achieved an accuracy comparable to ours that is 65% and 71% for two classifiers. However, they classified between two classes (finger and wrist movement).
Lehtonen et al. achieved a higher accuracy of 80% and 86% respectively. However, their chosen movements were of left and right hand. These movements have a larger spatial difference in motor cortex and hence, the classifier works better. They used common spatial subspace for feature extraction and support vector clustering as a classifier. Table 1 compares all the researches.
The main contribution of our research is to be able to differ between movements that can be used to control prosthesis. Since this study is focusing on the right hand only, the accuracy achieved between our four classes is higher than expected and the proposed method can be used to control prosthesis through EEG signals of the right hand finger movements.
Conclusion
It has been shown that PSD and logistic regression model can be used for classification of finger movements. A mean accuracy of 65% has been achieved, which are significantly improved results. This can be increased further if the overlapping classes of fist, index and middle finger are classified better and this is the suggested future work for this project. Comparison with other classifiers shows that logistic regression model performs best with the data set under consideration. Comparing research with other work has also shown that the accuracy achieved is greater than that was expected by these movements.
In future, the model is to be implemented on an embedded system, which will be then connected with prosthesis. This is will create a working model. The embedded system will then be modified to intake real-time data from the electrode headset. At the moment, work is being done on development of the embedded system of this model. The trained logistic regression classifier of the current research will be used. The filter and feature extractor will be developed in the embedded system. The parameters of the filter and PSD model will remain the same as the current research.
Once the system has been developed, it will be connected with upper limb prosthesis. Initially this will be controlled using the data that was collected for this study. The data will be sent to the controller to first create an offline system on an embedded system. In the next step of the study, the system can be made online with collection of data from the subject at the time of processing.
The research can be further refined by taking data from multiple volunteers rather than one and re-training the classifier according to that. This will result in a decrease in the accuracy of the classifier due to the different potentials of different humans. However, principal component analysis and other techniques can be used to improve that. In conclusion, the current research is a stepping stone for further research in the proposed movements and provides a baseline for future research.
References
- Jonathan RW, Niels B, Dennis J, Gert P, Theresa MV. Brain-computer interfaces for communication and control. Clin Neurophysiol 2002; 113: 767-791.
- Elad YT, Gideon FI. Feature selection for the classification of movements from single movement-related potentials. IEEE Trans Neural Systems Rehabil Eng 2002; 10.
- Dennis J, Laurie AM, Theresa MV, Jonathan RW. Mu and beta rhythm topographies during motor imagery and actual movements. Brain Topogr 2000; 12.
- Lotte F, Congedo M, Ecuyer AL, Lamarche F, Arnaldi B. Review of classification algorithms for EEG-based Brain-Computer Interfaces. J Neural Eng 2007.
- Ke L, Ran X, Jania G, Lei D. Decoding individual finger movements from one hand using human EEG signals. 2014.
- Ran X, Lei D. Evaluation of EEG features in decoding individual finger movements from one hand. Comput Mathematical Methods Med 2013.
- Vuckovic A, Sepulveda F. A four-class BCI based on motor imagination of the right and the left hand wrist. Appl Sci Biomed Commun Technol 2008.
- Mohamed AK, Marwala T, John LR. Single-trial EEG discrimination between wrist and finger movement imagery and execution in a sensorimotor BCI. EMBC Annual Int Conf 2011.
- Boyu W, Feng Wan. Classification of single-trial EEG based on support vector clustering during finger movement. 6th International symposium on neural networks, ISNN 2009.
- Lehtonen J, Jylänki P, Kauhanen L, Sams M. Online classification of single EEG trials during finger movements. IEEE Trans Biomed Eng 2008; 55.
- David WH, Stanley L, Rodney X. Applied logistic regression. 2013.
- Prabhu KMM. Window functions and their applications in signal processing. 2013.
- Nadia N, Luiza de MM, Springer. Evolvable machines: theory and practice. 2004.
- Ion M, Alexander Z. Bioinformatics research and applications: Third International symposium. ISBRA 2007.
- Christoph G, Alois S, Christa N, Dirk W, Thomas S, Gert P. Rapid prototyping of an EEG-Based Brain-Computer Interface (BCI). IEEE Trans Neural Systems Rehabil Eng 2001; 9.
- Kalaivani M, Kalaivani V, Anusuya DV. Analysis of EEG signals for the detection of brain abnormalities. ICSCN 2014.
- Vučković A, Sepulveda F. A two stage four class BCI based on imaginary movements of the left and right wrist. 2012.
- Yong L, Xiaorong G, Hesheng L, Shangkai G. Classification Of single-trial electroencephalogram during finger movement. IEEE Trans Biomed Eng 2004; 51.
- Bernardo DS, Matteo M, Luca TM. Online detection of P300 and error potentials in a BCI speller. J Comput Intell Neurosci 2010.